Lindholm Chapter 10 - Generative Models and Unlabelled Data
Lindholm Chapter 10 - Generative Models and Unlabelled Data
Gaussian Mixture Model and Discriminant Analysis.
- The Gaussian Mixture Model makes use of the factorisation of the joint PDF
- The second factor is the marginal distribution of .
- Since is categorical and thereby takes values in the set , this is given by a categorical distribution with parameters, .
- The first factor in Equation 10.1a is the class-conditional distribution of the input for a certain class . In a classification setting, it is natural to assume that these distributions are different for different classes of .
- If it is possible to predict the class based on the information in then the characteristics (distribution) of should depend on .
- Need to make further assumptions on these class-conditional distributions.
- The basic assumption for a GMM is that each is a Gaussian Distribution with a class-dependent mean vector and covariance matrix .
Supervised Learning of the Gaussian Mixture Model
- The unknown parameters to be learned are .
- Start with the supervised case, meaning that the training data contains inputs , and corresponding outputs - .
- Mathematically we learn the GMM by maximising the log-likelihood of the training data:
- Due to the generative model, this is based on the joint likelihood of the inputs and outputs.
- From this, we derive the model definition is given by:
- The indicator function effectively separates the log-likelihood function into independent sums - one for each class
- Separates based on the class labels of the training points.
- The optimisation problem has a close-form solution.
- Begin with marginal class probabilities, , where is the number of training points in class .
- Therefore and thus .
- The mean vector of each class is estimated as:
- The equations 10.3b and 10.3c learns a Gaussian distribution for for each class such that the mean and covariance matching.
- Note that can be computed regardless of whether the data really comes from a Gaussian distribution or not.
Predicting Output Labels for New Inputs: Discriminant Analysis
- The key insight for using generative model to make predictions is to realise that predicting the output for a know value amounts to computing the conditional distribution :
- From this, we get the classifier:
- We obtain the hard predictions by selecting the class with the highest probability prediction
- Taking the logarithm and noting that only the numerator in Equation 10.5 depends on , we can equivalently write:
- Since the logarithm of the Gaussian PDF is a quadratic function in , the decision boundary for this classifier is quadratic - this is called Quadratic Discriminant Analysis (QDA).
- If we make an additional simplifying assumption about the model, instead Linear Discriminant Analysis (LDA) is obtained.
- The simplifying assumption is that the covariance matrix is equal for all classes, i.e. for all .
- This new covariance matrix is replaced by the covariance matrix learned by the following equation.
- This simplifying assumption results in a cancellation of a cancellation of all quadratic terms when computing class probabilities, when computing class predictions using Equation 10.7.
- Consequently, LDA is a linear classifier just like Logistic regression - will often perform similarly.
- Will not perform identically as parameters are learned in different ways.
- The pseudocode for training the GMM is given as:
Data Training data .
Result Gaussian Mixture Model
- for do
- | Compute using Equation 10.3a, using Equation 10.3b and using Equation 10.3c
- end
- The pseudocode for predicting using the GMM is given as:
Data Gaussian Mixture Model and test input .
Result Prediction .
- for do
- | .
- end
- Set
Semi-Supervised Learning of the Gaussian Mixture Model
Want to exploit the information available in the unlabelled data to end up with a better model
Consider semi-supervised learning problem, where some of the output values are missing in the training data.
The input values for which the output is missing are called unlabelled data points.
Denote the total number of training points as , out of which only are labelled input-output pairs and the remaining unlabelled data points are .
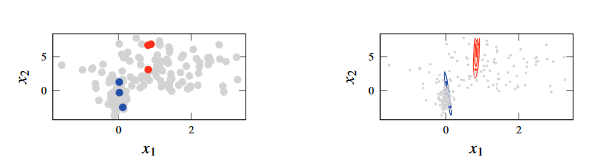
Figure 1 - Semi-supervised learning, in which which we have fitted a GMM to data in which we do not have most of the labels. The unlabelled data has evidently made the problem harder
One way to approach a semi-supervised problem is to use a generative model.
A generative model is a model of the joint distribution which can be factorised as .
- Since the marginal distribution of the inputs is a part of the model, it seems plausible that the unlabelled data can be useful.
Intuitively, the unlabelled inputs can be used to find groups of input values with similar properties - which are then assumed to be of the same class.
Use the maximum-likelihood approach - seek model parameters that maximise likelihood of observed data.
- This problem has no closed-form solution.
- When computing the mean vector for the th class we do not know which points should be included in the sum.
- We could first learn an initial GMM which is then used to predict the missing labels.
- We then use these predictions to update the model.
- This iterative process results in te following algorithm:
- Learn the GMM from the labelled input-output pairs
- Use the GMM to predict (as a QDA classifier) the missing outputs
- Update the GMM including the predicted outputs from step 2
- Repeat steps (2) and (3) until convergence
- From Step 2, we should return the predicted class probabilities and not the class predictions predicted using the current parameter estimates
- We make use of the predicted class probabilities using the Equation 10.10a
- We update the parameters using the following equations
- The pseudocode for the semi-supervised learning of the GMM is given as:
Data: Partially labelled training data with output classes .
Result Gaussian Mixture Model
- Compute according to Equation 10,3 using only the labelled data
- repeat
- | For each in the unlabelled subset, compute the prediction according to Equation 10.5 using the current parameter estimates
- | Update the parameter estimates according to Equation 10.10
- until convergence
- The prediction is done in an identical fashion to QDA (Method 10.1)
Cluster Analysis
Models in supervised learning have the objective to learn input-output relationship based on examples.
In semi-supervised learning, mixed labelled and unlabelled data to learn a mode which makes use of both sources of information
In unsupervised learning assume that all points are unlabelled.
Objective is to build a model that can be used to reason about key properties of the data (or the process usd to generate the data)
Clustering is highly related to classification
Assign a discrete index to each cluster and say all values in the th cluster are of class .
- Difference between clustering and classification is that we train based on the values without any labels.
Unsupervised Learning of the Gaussian Mixture Model
- The GMM is a joint model for and , given by
- To obtain a model for only we can marginalise out from it.
- That is, we consider as being a latent random variable - a random variable that exists in the model but is not observed in the data.
- In practice, still lean the joined model, but from data containing only values.
- We want to learn which values come from the same class-conditional distribution based on their similarity.
- That is, the latent variables need to be inferred from this data, and then use this latent data to fit the model parameters.
- We can modify Method 10.2 for semi-supervised learning to work with completely unlabelled data - need to replace the initial weight initialisation.
- The pseudocode for the unsupervised learning of the GMM is given as:
Data Unlabelled training data , number of clusters .
Result Gaussian Mixture Model
- Initialise
- repeat
- | For each in compute the prediction according to Equation 10.5 using the current parameter estimates
- | Update the parameter estimates according to Equation 10.16
- This is the EM algorithm - a tool for solving maximum likelihood problems with latent variables.
- In the “E step” in line 3, we compute the responsibility values - how likely it is that a given data point could have been generated by each of the components in the mixture models.
- In the “M step” in line 4, we update the parameters of the mixture model based on the responsibility values.
- In the “E step” in line 3, we compute the responsibility values - how likely it is that a given data point could have been generated by each of the components in the mixture models.
- The step can be summarised as updating the parameters as follows:
k-Means Clustering
- Alternative