COMP4702 Lecture 11
COMP4702 Lecture 11
Chapter 10: Generative Models
- Previous chapters have been mostly about supervised learning, which generate models of .
- Generative models are models of .
- Dropping the term would make this unsupervised learning.
- Generative models are very similar to probabilistic models and computer simulation models - they all draw from some probability distribution.
- So we can use a probability density estimation and think of this as a foundation for unsupervised learning and generative models.
Gaussian Mixture Models and Discriminant Analysis
Mixture Densities
- A mixture model is a weighted sum of component densities e.g. this case, Gaussian Distributions
- Where is the th component (aka group or cluster assumed to be in the data)
- is the prior/mixture coefficient or weight of the th component and is the th component PDF.
- Note that and .
- We will look at Gaussian Mixture Models (GMMs), so:
- If for a single variable, specify mean and standard deviation.
- If multivariate, specify mean vector and covariance matrix.
- The parameters of our GMM are . So the LHS of the above equation is more properly written as:
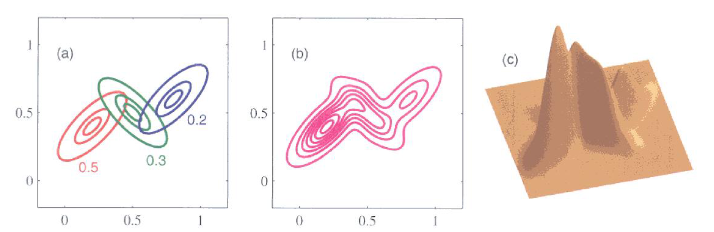
Figure 1 - A Gaussian mixture model with 3 Gaussian distributions in a two-dimensional space.
- The left plot Figure 1 describes that we can write a GMM as the weighted sum of 3 (in this case) Gaussian distributions.
- These three smooth functions add up to create a single smooth function as shown in the center plot of Figure 1.
- This GMM is plotted in three dimensions in the right of Figure 1.
- What happens next is really all about whether or not you know apriori whether your data comes from some groups/categories/classes and if you have those “labels” for your data.
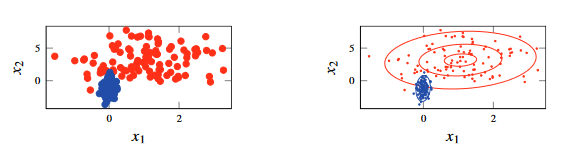
Figure 2 - A Gaussian mixture model with 3 Gaussian distributions in a two-dimensional space.
- The above example assumes that you know the labels , shown on the left.
- With these knowledge, you can separate these points out and create a Gaussian distribution for each class - shown by the contour lines in the plot to the right.
- The weight of each probability distributed by the frequency of each class in the data.
- Can use Maximum Likelihood Estimation (MLE) to estimate the parameters of each Gaussian distribution.
Predicting Output Labels for New Inputs: Discriminant Analysis
This section shows that you can use a GMM to do classification, because you can use your model of to get .
- being the coordinates and being the class labels.
This is a nice illustration for multi-class classification, bu you can just use a single Gaussian distribution for the binary classification and you’d be close to logistic regression
However, there are a few key differences:
Priors: not present in logistic regression. With Logistic Regression, it doesn’t explicitly encode the idea that one class has more samples than the other.
Different parametrisations of the covariance matrix give rise to Linear Discriminant Analysis (LDA) or Quadratic Discriminant Analysis (QDA) when you allow covariance modelling.
- LDA when you have a diagonal Covariance matrix.
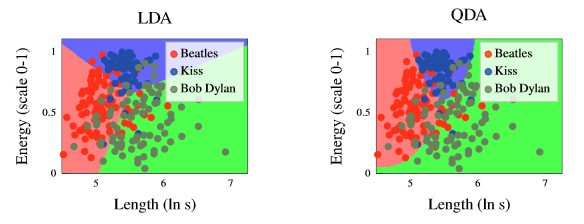
Figure 3 - Difference in decision boundaries generated by LDAs and QDAs.
The figure above shows the difference between the decision boundaries generated by Linear Discriminant Analysis (LDA) vs Quadratic Discriminant Analysis (QDA).
- Observe that in the figure to the left (LDA), the decision boundaries are linear whereas the decision boundaries in the figure to the right (QDA) are quadratic.
Semi-Supervised Learning of the Gaussian Mixture Model.
- Where we have some data that is labelled, but also (probably a lot more) that is unlabelled.
- We would like to make use of this unlabelled data as well as the labelled data.
- We would like to maximise the likelihood but we can’t do this directly using the unlabelled data as shown in Equation 10.9:
- However, this problem has no closed-form solution
- However, we can take the following approach:
- Learn the GMM from he labelled data.
- Use the GMM to predict as a proxy for the labels for the unlabelled data.
- Update the GMM’s parameters including the data with predicted labels.
- This concept works and isn’t just some random idea - it’s a version of the Expectation Maximisation (EM) algorithm
- The equations relevant to this procedure are given as:
Cluster Analysis
If we do not have a defined target variable (e.g., labels or numerical variables) we are in the realm of unsupervised learning.
The data consists of observations of features collected from some problem domain, and presumably contains structure and information that is of interest.
One way to learn about that structure is to build a model of the distribution of the data (i.e. probability density estimation).
A GMM is a flexible probability density estimator, and peaks of the density give us an understanding of the clustering of data (places in the feature space where data occurs with high probability).
The EM algorithm can be used to fit a GMM to data for .
- Effectively is marginalised out of the estimate and becomes a latent variable for the model.
- The latent variables are where is the cluster index for data point .
- This algorithm is shown in Method 10.3 below:
Data Unlabelled training data , number of clusters Result Gaussian Mixture Model
- Initialise
- repeat
- | For each in compute the prediction using Equation 10.5 using the current parameter estimate .
- | Update the parameter estimates using Equations 10.10.
- until convergence
In the “E step” (Line 3) We compute the “responsibility values” - how likely is it that a given data point could have been generated by each of the components of the mixture model? Note that the notation here is quite confusing: is actually a vector of values for each mixture component.
In the “M step” (Line 4) we update the model parameters using the responsibility values computed in the E step. Lindholm refer to the responsibility values as weights,
Maximising the likelihood for a GMM is a non-convex optimisation problem, and the EM algorithm performs local optimisation which hopefully converges toward a stationary point
This works pretty well most of the time, though there is also a degeneracy in the problem - undesirable solutions where the likelihood diverges to infinity.
k-Means Clustering
- We can understand that the -Means clustering algorithm as a simplified version of fitting a GMM using the EM algorithm.
- The simplification made are:
- Rather than calculating (soft, probabilistic) responsibility values, -Means calculates “hard” responsibility values based on the distances between a point and a set of cluster centers.
- The distances in the GMM use the covariance information (i.e. the Mahalanobis distance between each data point and each component of the GMM)
- The algorithm is given as the following steps:
- Set the cluster centers to some initial values
- Determine which cluster each belongs to by finding the cluster center that is for all .$
- Update the cluster centers as the average of all that belongs to
- Repeat steps 2 and 3 until convergence
Choosing the Number of Clusters
- Unfortunately no simple answer - sometimes problem domain knowledge can guide the choice.